A general implementation of Structural Equation Models with latent variables (MLE, 2SLS, and composite likelihood estimators) with both continuous, censored, and ordinal outcomes (Holst and Budtz-Joergensen (2013) <10.1007/s00180-012-0344-y>). The package also provides methods for graph exploration (d-separation, back-door criterion), simulation of general non-linear latent variable models, and estimation of influence functions for a broad range of statistical models.
For graphical capabilities the Rgraphviz package is needed (first install the BiocManager package)
or the igraph or visNetwork packages
The development version of lava may also be installed directly from github:
To cite that lava package please use one of the following references
Klaus K. Holst and Esben Budtz-Joergensen (2013). Linear Latent Variable Models: The lava-package. Computational Statistics 28 (4), pp 1385-1453. http://dx.doi.org/10.1007/s00180-012-0344-y
@article{lava,
title = {Linear Latent Variable Models: The lava-package},
author = {Klaus Kähler Holst and Esben Budtz-Jørgensen},
year = {2013},
volume = {28},
number = {4},
pages = {1385-1452},
journal = {Computational Statistics},
doi = {10.1007/s00180-012-0344-y}
}Klaus K. Holst and Esben Budtz-Jørgensen (2020). A two-stage estimation procedure for non-linear structural equation models. Biostatistics 21 (4), pp 676-691. http://dx.doi.org/10.1093/biostatistics/kxy082
@article{lava_nlin,
title = {A two-stage estimation procedure for non-linear structural equation models},
author = {Klaus Kähler Holst and Esben Budtz-Jørgensen},
journal = {Biostatistics},
year = {2020},
volume = {21},
number = {4},
pages = {676-691},
doi = {10.1093/biostatistics/kxy082},
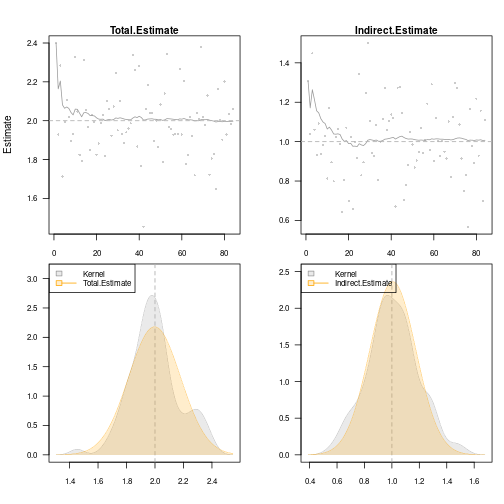
}Specify structural equation models with two factors
m <- lvm()
regression(m) <- y1 + y2 + y3 ~ eta1
regression(m) <- z1 + z2 + z3 ~ eta2
latent(m) <- ~ eta1 + eta2
regression(m) <- eta2 ~ eta1 + x
regression(m) <- eta1 ~ x
labels(m) <- c(eta1=expression(eta[1]), eta2=expression(eta[2]))
plot(m)
Simulation
Estimation
e <- estimate(m, d)
e
#> Estimate Std. Error Z-value P-value
#> Measurements:
#> y2~eta1 0.95462 0.08083 11.80993 <1e-12
#> y3~eta1 0.98476 0.08922 11.03722 <1e-12
#> z2~eta2 0.97038 0.05368 18.07714 <1e-12
#> z3~eta2 0.95608 0.05643 16.94182 <1e-12
#> Regressions:
#> eta1~x 1.24587 0.11486 10.84694 <1e-12
#> eta2~eta1 0.95608 0.18008 5.30910 1.102e-07
#> eta2~x 1.11495 0.25228 4.41951 9.893e-06
#> Intercepts:
#> y2 -0.13896 0.12458 -1.11537 0.2647
#> y3 -0.07661 0.13869 -0.55241 0.5807
#> eta1 0.15801 0.12780 1.23644 0.2163
#> z2 -0.00441 0.14858 -0.02969 0.9763
#> z3 -0.15900 0.15731 -1.01076 0.3121
#> eta2 -0.14143 0.18380 -0.76949 0.4416
#> Residual Variances:
#> y1 0.69684 0.14858 4.69004
#> y2 0.89804 0.16630 5.40026
#> y3 1.22456 0.21182 5.78109
#> eta1 0.93620 0.19623 4.77084
#> z1 1.41422 0.26259 5.38570
#> z2 0.87569 0.19463 4.49934
#> z3 1.18155 0.22640 5.21883
#> eta2 1.24430 0.28992 4.29195Assessing goodness-of-fit, here the linearity between eta2 and eta1 (requires the gof package which can installed from CRAN)
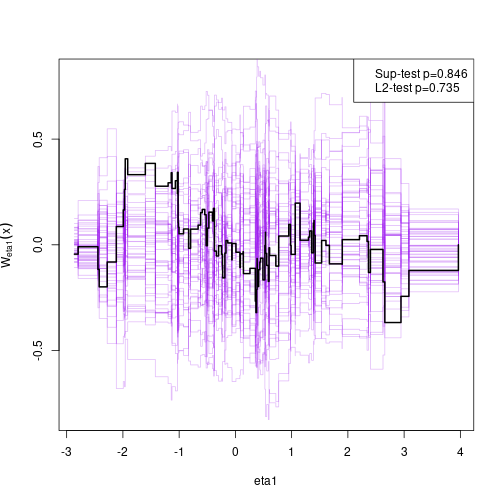
Simulate non-linear model
m <- lvm(y1 + y2 + y3 ~ u, u ~ x)
transform(m,u2 ~ u) <- function(x) x^2
regression(m) <- z~u2+u
d <- sim(m,200,p=c("z"=-1, "z~u2"=-0.5), seed=1)Stage 1:
m1 <- lvm(c(y1[0:s], y2[0:s], y3[0:s]) ~ 1*u, u ~ x)
latent(m1) <- ~ u
(e1 <- estimate(m1, d))
#> Estimate Std. Error Z-value P-value
#> Regressions:
#> u~x 1.06998 0.08208 13.03542 <1e-12
#> Intercepts:
#> u -0.08871 0.08753 -1.01344 0.3108
#> Residual Variances:
#> y1 1.00054 0.07075 14.14214
#> u 1.19873 0.15503 7.73233Stage 2
pp <- function(mu,var,data,...) cbind(u=mu[,"u"], u2=mu[,"u"]^2+var["u","u"])
(e <- measurement.error(e1, z~1+x, data=d, predictfun=pp))
#> Estimate Std.Err 2.5% 97.5% P-value
#> (Intercept) -1.1181 0.13795 -1.3885 -0.8477 5.273e-16
#> x -0.0537 0.13213 -0.3127 0.2053 6.844e-01
#> u 1.0039 0.11504 0.7785 1.2294 2.609e-18
#> u2 -0.4718 0.05213 -0.5740 -0.3697 1.410e-19f <- function(p) p[1]+p["u"]*u+p["u2"]*u^2
u <- seq(-1, 1, length.out=100)
plot(e, f, data=data.frame(u))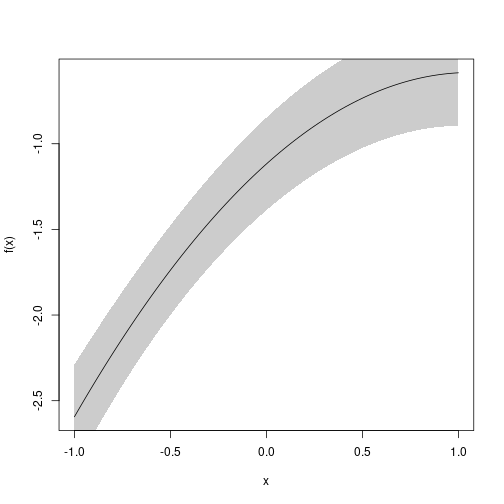
Studying the small-sample properties of a mediation analysis
m <- lvm(y~x, c~1)
regression(m) <- y+x ~ z
eventTime(m) <- t~min(y=1, c=0)
transform(m,S~t+status) <- function(x) survival::Surv(x[,1],x[,2])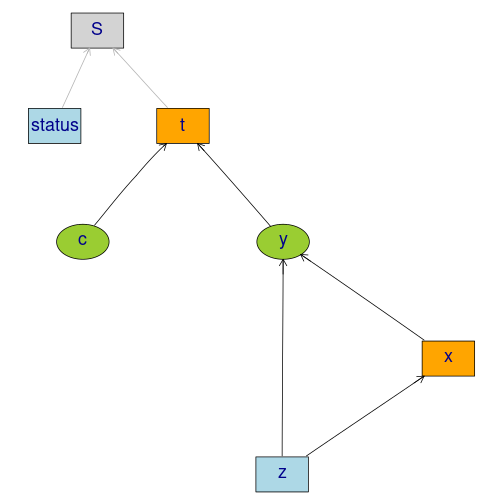
Simulate from model and estimate indirect effects
onerun <- function(...) {
d <- sim(m, 100)
m0 <- lvm(S~x+z, x~z)
e <- estimate(m0, d, estimator="glm")
vec(summary(effects(e, S~z))$coef[,1:2])
}
val <- sim(onerun, 100)
summary(val, estimate=1:4, se=5:8, short=TRUE)
#> 100 replications Time: 3.667s
#>
#> Total.Estimate Direct.Estimate Indirect.Estimate S~x~z.Estimate
#> Mean 1.97292 0.96537 1.00755 1.00755
#> SD 0.16900 0.18782 0.15924 0.15924
#> SE 0.18665 0.18090 0.16431 0.16431
#> SE/SD 1.10446 0.96315 1.03183 1.03183
#>
#> Min 1.47243 0.54497 0.54554 0.54554
#> 2.5% 1.63496 0.61228 0.64914 0.64914
#> 50% 1.95574 0.97154 0.99120 0.99120
#> 97.5% 2.27887 1.32443 1.27807 1.27807
#> Max 2.45746 1.49491 1.33446 1.33446
#>
#> Missing 0.00000 0.00000 0.00000 0.00000Add additional simulations and visualize results
val <- sim(val,500) ## Add 500 simulations
plot(val, estimate=c("Total.Estimate", "Indirect.Estimate"),
true=c(2, 1), se=c("Total.Std.Err", "Indirect.Std.Err"),
scatter.plot=TRUE)